What are we interested in?
Electron transfer (ET) reactions unify many energy transduction processes in chemistry and biochemistry. Research in the Warren group focuses on understanding ET reactions in protein model systems and in electrocatalysts. We have focused interests in how microenvironments affect redox reactions.
Electron Flow in Protein Structures
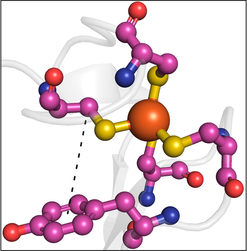
Our work on ET in proteins involves aromatic amino acids, especially tyrosine and tryptophan. Surveys of structural data revealed interesting associations between tyrosine and methionine. Ongoing work involves building a number of protein models to explore these interesting associations via ET chemistry.
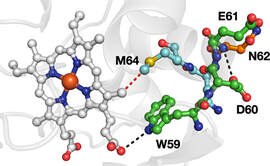
In other work on protein ET, we are investigating how protein microenvironment affects long-range ET in photosensitizer-modified proteins. Studies of artificial tyrosine derivatives enable detailed optical and magnetic resonance spectroscopies.
Small Molecule Electrocatalysis
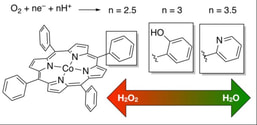
Some of our work on electrocatalysts involves the investigation of different ancillary groups on metalloporphyrins, from hydrogen bond promoters to local electrostatic interactions, and how they affect the selectivity and kinetics of oxygen and carbon dioxide reduction.
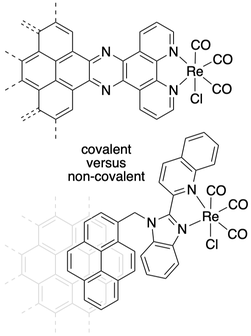
Our other research efforts involve exploration of differences in heterogeneous and homogeneous carbon dioxide chemistry, for example using the rhenium(I) complexes with specific ligand designs to attach to electrode surfaces.
Our other research efforts involve exploration of differences in heterogeneous and homogeneous carbon dioxide chemistry, for example using the rhenium(I) complexes with specific ligand designs to attach to electrode surfaces.
